reduced chi homepage
0.14
News
- Date:
- Fri Aug 29 17:22:09 PDT 2008 0.14 tag release
Fri Oct 19 15:31:22 PDT 2007 0.13 tag release
Mon May 7 00:42:01 PDT 2007 0.12 tag release
Thu Mar 22 01:21:37 PDT 2007 0.11 tag release
Thu Jun 1 20:27:03 PDT 2006 0.10.1 tag release
Wed May 17 02:49:44 PDT 2006 Initial release!
About
Reduced chi is a simple crystallographic program used to calculate the "reduced chi" value for a dataset:
![\[ \frac{\chi}{\nu} = \sqrt{\frac{\sum\limits_\mathrm{hkl}^{\mathrm{n}} {\frac{(F_{\mathrm{obs}}-\sigma_{\mathrm{A}}F_{\mathrm{calc}})^2} {\sigma_{F_{\mathrm{obs}}}^2+\sigma_{\Delta}^2}}} {(n-p_D)}} \]](form_0.png)
this is done using a mix of C and C++, much of which is simplified thanks to several high level libraries. Other calculated values include the R, Rfree, and the signal to noise ratio of the data. It also serves as an excellent platform for experimentation, as the source code is small, simple and quite adaptable.
As of version 0.11, redchi also implements a refinement routine using a maximum likelihood principle:
![\[ -\mathrm{log}(L(\theta)) = -\mathrm{log}(\mathrm{Pr}_\mathrm{geom}|\theta) - \mathrm{log}(\mathrm{Pr}_\mathrm{xray}|\theta) \]](form_1.png)
the program also computes the Deviance information criterion:  for a given refinement, a measure of Bayesian "goodness" of fit. In this equation,
for a given refinement, a measure of Bayesian "goodness" of fit. In this equation,  represents the effective number of parameters, or more precisely, the difference in the posterior mean of the deviance (where deviance is measured as
represents the effective number of parameters, or more precisely, the difference in the posterior mean of the deviance (where deviance is measured as  ) and the deviance at the posterior means (i.e.
) and the deviance at the posterior means (i.e. 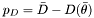 ). This can be used in the reduced chi calculation above in order to compute the number of degrees of freedom.
). This can be used in the reduced chi calculation above in order to compute the number of degrees of freedom.
However, the refinement methods used in redchi are still under development, so please send any error/bug reports to the authors.
Basically, redchi takes a command file as input (see Input file commands), parses it and generates text output. It can optionally generate 2D graphs of your data as well, thanks to Grace. To run redchi, you'll need an MTZ file containing observed structure factors and errors (optionally containing Rfree flagged reflections), and a PDB file. Thats it! To get started, check out some Examples.
redchi manual:
- Command line options
- Input file commands
- Installation/obtaining
- Examples
- Sourceforge home for redchi
- Changelog
License and Availability
redchi is licensed under the GNU General Public License version 3 (GPLv3), and is available free of charge in both binary and source form.
 1.5.6
1.5.6